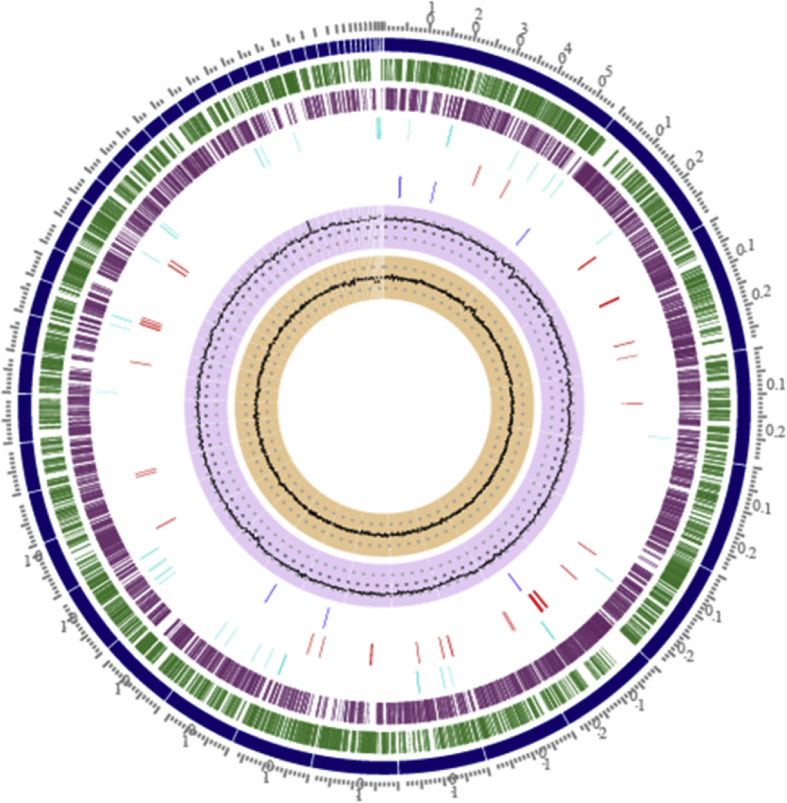
The bacterial isolates of genus Rhodococcus are greatest recognized for his or her important biodegradation skills. Here, we report the information associated to draft genome sequencing of Rhodococcus rhodochrous strain SPC17 isolated from sediments of Lonar Lake.
The de novo meeting of 1598096 Illumina’s paired-end sequencing reads resulted in 51 contigs for an general genome meeting dimension of 4.98Mb.
A complete of 4546 genes have been predicted utilizing the National Center for Biotechnology Information- Prokaryotic Genome Annotation Pipeline (NCBI-PGAP). RAST server-based annotation of the Rhodococcus strain SPC17 genome resulted in a complete of 295 subsystems with 25% subsystem protection.
The information on the draft genome shotgun mission are accessible at NCBI-GenBank below the accession quantity WUUR00000000. Our information useful resource will facilitate additional molecular and genomic research of various hydrocarbon catabolizing genes current in Rhodococcus rhodochrous strain SPC17.
Comprehensive annotations of human herpesvirus 6A and 6B genomes reveal novel and conserved genomic options.
Human herpesvirus-6 (HHV-6) A and B are ubiquitous betaherpesviruses, infecting the bulk of the human inhabitants. They embody giant genomes and our understanding of their protein coding potential is way from full.
Here, we make use of ribosome-profiling and systematic transcript-analysis to experimentally outline HHV-6 translation merchandise. We determine a whole bunch of new open studying frames (ORFs), together with upstream ORFs (uORFs) and inside ORFs (iORFs), producing a whole unbiased atlas of HHV-6 proteome.
By integrating systematic information from the prototypic betaherpesvirus, human cytomegalovirus, we uncover quite a few uORFs and iORFs conserved throughout betaherpesviruses and we present uORFs are enriched in late viral genes.
We recognized three extremely ample HHV-6 encoded lengthy non-coding RNAs, one of which generates a non-polyadenylated steady intron showing to be a conserved characteristic of betaherpesviruses. Overall, our work reveals the complexity of HHV-6 genomes and highlights novel options conserved between betaherpesviruses, offering a wealthy useful resource for future useful research.
MicroScope: an built-in platform for the annotation and exploration of microbial gene capabilities by way of genomic, pangenomic and metabolic comparative analysis.
Large-scale genome sequencing and the more and more huge use of high-throughput approaches produce an enormous quantity of new info that fully transforms our understanding of 1000’s of microbial species.
However, regardless of the event of highly effective bioinformatics approaches, full interpretation of the content material of these genomes stays a tough process.
Here we current new enhancements of the MicroScope consumer interface for genome choice, navigation and knowledgeable gene annotation. Automatic useful annotation procedures of the platform have additionally been up to date and we added a number of new instruments for the useful annotation of genes and genomic areas.
We lastly focus on new instruments and pipeline developed to carry out comparative analyses on a whole bunch of genomes based mostly on pangenome graphs. To date, MicroScope comprises information for >11 800 microbial genomes, half of that are manually curated and maintained by microbiologists (>4500 private accounts in September 2019). The platform allows collaborative work in a wealthy comparative genomic context and improves community-based curation efforts.
